The ibidi µ-Patterning Technology
Achieve Precise Spatial Control Over Cell Adhesion With ibidi Micropatterning
The ibidi µ-Patterning technology enables spatially defined cell adhesion for various 2D and 3D cell culture applications.
Miniaturized adhesive patterns (e.g., squares or dots) are irreversibly printed on the non-adhesive Bioinert surface of the ibidi Polymer Coverslip, allowing for precisely controlled cell adhesion. The µ-Patterns are dry-stable, sterile, and ready to use.
The µ-Pattern, the Bioinert surface, and the ibidi Polymer Coverslip are all optimized for high resolution imaging and microscopy.
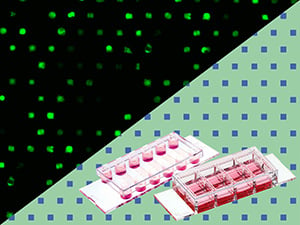
ibidi's Ready-to-Use µ-Patterning Products
Understanding the Key Elements of µ-Patterns
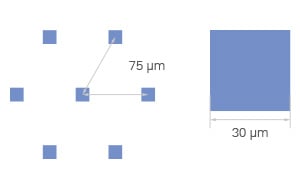
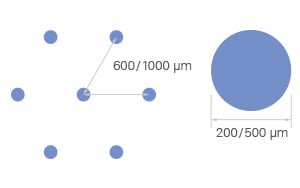
Every ibidi µ-Pattern is designed with precise spatial control, ensuring reproducible cell adhesion and organization. Let's break down the essential pattern components:
- Shape – Defines the geometry of the attachment area (e.g., cir = circle)
- Size – The diameter or width of the attachment area, measured in micrometers (µm) (e.g., 500 µm)
- Pitch – The spacing between adjacent shapes, measured from center to center (e.g., pit1000 = 1000 µm)
- Layout – The overall pattern arrangement, determining how attachment areas are structured (e.g., a hexagonal layout)
- Surface – The cell attachment surface where cells adhere, made of ibiTreat for optimal cell growth
Application Examples
Single-Cell Arrays
The size of the µ-Pattern can be adapted to the morphology of the cell type of interest, so that an array of single cells can be conveniently analyzed using applications such as high-resolution imaging.
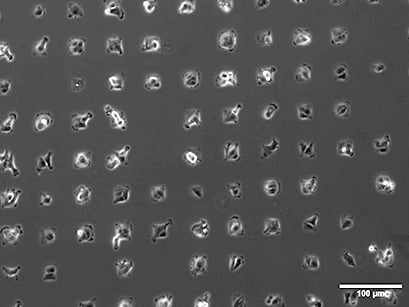

Single-cell array with RCC26 tumor cells. Spot size 40 µm x 40 µm. Phase contrast microscopy, 10x objective lens.
Spheroid Generation
Defined adhesion spots, surrounded by Bioinert, are able to catch all adherent single cells from a cell suspension. Bioinert is fully non-cell-attachable. This forces all cells to aggregate to each other at the adhesion spots, thus forming spheroids in a defined and controllable way.
Suspension of NIH-3T3 cell line seeded on 200 µm adhesion spots, 64 hours live cell imaging, phase contrast, 4x objective lens.
Interested in unlocking new possibilities with ibidi µ-Patterning?
Let's collaborate on a custom solution to meet your research needs! We will work with you to co-develop a tailor-made design here.

You need support or want to test the ibiTreat µ-Patterns first? Contact us at: info@ibidi.com or choose up to three free samples from our ibiTreat micropatterning standard products!
The size and the cell adhesion properties of the µ-Patterning can be adjusted to hold 3D spheroids and microtissues in position. Using this setup, cells in 3D can be studied during proliferation, differentiation, invasion, and migration.
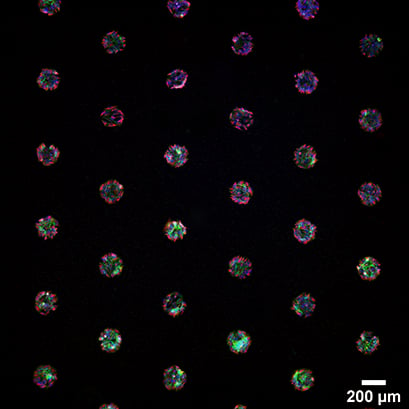
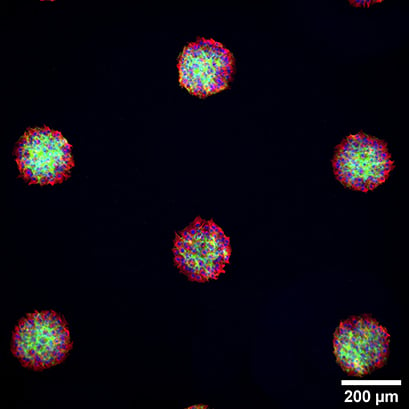
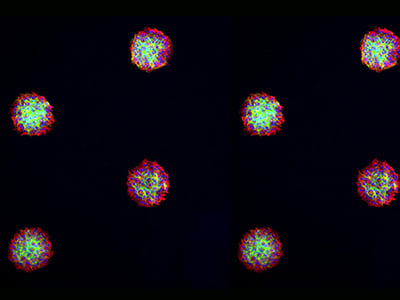
Immunofluorescence staining of spheroids from RCC26 (left) and Rat1 cells (right) in in the µ-Slide VI 0.4 Luer, patterned with 200 µm circles at 600 µm distance. The cells were stained with phalloidin (green) and alpha-tubulin (red). Nuclei were stained with DAPI (blue). Widefield fluorescence microscopy, 4x and 10x objective.
Line Arrays
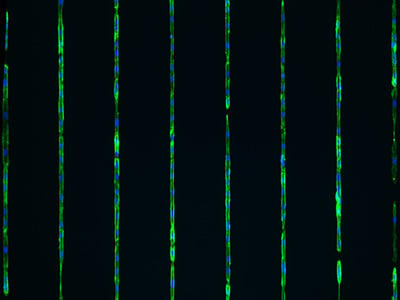
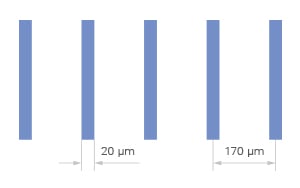
The ready-to-use structured microlane environment supports e.g., neuronal cell and organelle outgrowth, including neurite extension and cell–cell communication. They provide a platform for studying geometry-driven cell migration and polarization, cytoskeletal dynamics, and organelle transport.
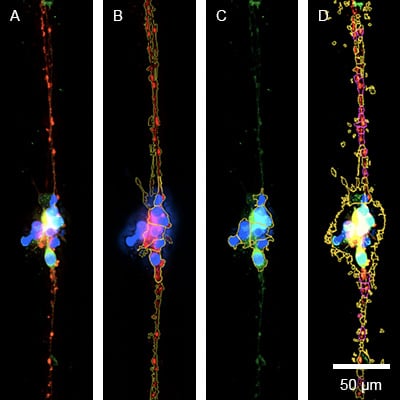
Neuronite outgrowth analysis. Fluorescence overlay image (A) of ß3 tubulin (red), VDAC (Mitochondria) (green/yellow), and the cell nuclei (DAPI). Visual segmentation is indicated by yellow lines to measure the ß3 tubulin staining (B), the VDAC signal in soma (C), and the VDAC signal in axons (indicated in pink) (D). Human iPSC derived sensory neurons were incubated on the μ-Slide VI 0.4 μ-Pattern ibiTreat, lin20, pit170 for 5 days and imaged on a Cytation 7 (Agilent) with a 40x objective lens. Image courtesy of Moran Amit's Lab, University of Texas.
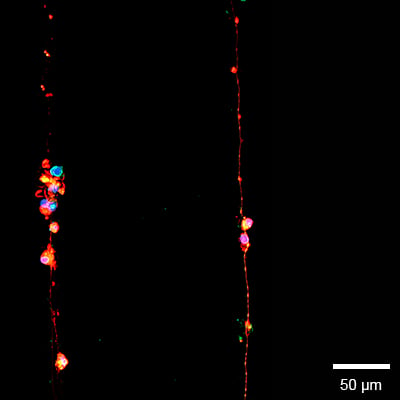
Voltage-dependent anion channel/mitochondrial porins (VDAC) and ß3-tubulin analysis to show the distribution of mitochondria along the cell. Fluorescence overlay image (A) of ß3-tubulin (red), VDAC (mitochondrial porins) (green/yellow) and the cell nuclei (DAPI). Human iPSC derived sensory neurons were incubated on the μ-Slide VI 0.4 μ-Pattern ibiTreat, lin20, pit170 for 5 days and imaged on a Cytation 7 (Agilent) with a 40x objective lens. Image courtesy of Moran Amit's Lab, University of Texas.
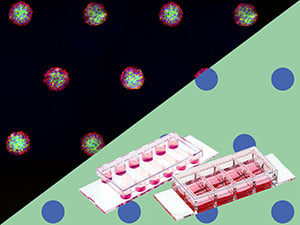
Integrating u-Patterning With the ibidi Pump System: Spheroid Culture Under Constant Media Circulation
By creating micropatterns within a µ-Slide VI 0.4 Luer, aggregates on the pattern can be constantly supplied with fresh media by perfusing the system with the ibidi Pump System.
By applying a constant media flux around the cell aggregates, spheroids become more compact and round.

3T3 fibroblasts were seeded on a patterned µ-Slide VI 0.4. A shear stress of 3 dyn/cm2 was applied 7 days after cell seeding. 15 h time lapse microscopy, 4x objective.
CAR-T Cell Killing Assays
CAR-T cells represent a promising new cancer therapy tool. Live cell imaging allows to analyze T cell/cancer cell interaction in real time with single cell resolution. However, analysis of confluent cell layers is very time-consuming and therefore not possible in high throughput screens. To facilitate high throughput label-free analysis of T cell potency in a live cell imaging setup, we generated arrays of homogenously distributed cancer cells. By combining optical analysis and advanced image processing, cytotoxic T cell activity over time on a single cell level can be evaluated without the use of any labeling.
Single Cells in a 2D Environment
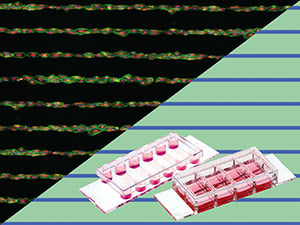
Time lapse microscopy of a CAR-T cell killing assay with RCC-26 tumor cells and JB4 T cells on a single cell pattern (µ-Slide VI 0.4, 20 µm squares, 120 µm distance, rectangular). Data were analyzed using FastTrack AI by MetaVi Labs.
Multi-Cell Spots in a 3D Collagen Matrix
RCC-26 cancer cells immobilized on multi-cell pads (µ-Slide VI 0.4, 200 µm circles, 600 µm distance, hexagonal). Effector T cells applied in a collagen I matrix (Collagen Type I, Rat Tail) induce apoptotic body formation of cancer cells.
Scientific Posters
Micropatterned Adhesion Sites for Spheroid Cultivation Under Flow (PDF)
Presented at the Annual Meeting of the Biophysical Society 2020, San Diego, USA.
Presented at the ASCB|EMBO Meeting 2019, Washington DC, USA.
Controlled Cell Adhesion With ibidi Micropatterning (PDF)
Presented at the µTAS Conference 2019, Basel, Switzerland.
Frequently Asked Questions
What is micropatterning?
Micropatterning is a technique used to control exactly where cells attach and grow on a surface. By defining adhesive regions with specific shapes and sizes, researchers can control cell position, spreading, and organization in a highly reproducible way. This helps create more standardized experiments and reduces variability in microscopy-based assays. Because the cellular microenvironment strongly influences behavior, precise spatial control can be critical for studying biological processes in a reproducible and quantitative manner.
What are the advantages of micropatterning?
Micropatterning helps you control where cells attach and how they are arranged on the culture surface. Instead of cells growing randomly, they grow in defined positions and shapes, making experiments easier to reproduce and compare. This can simplify microscopy, cell tracking, and quantitative image analysis, especially when cell shape, spacing, or cell–cell contact influences the biological outcome. ibidi provides ready-to-use µ-Patterning labware that can be integrated into standard cell culture workflows without requiring complex patterning equipment or specialized fabrication steps, making micropatterning accessible even for researchers who are new to the technique.
How does micropatterning work?
In general, micropatterning works by creating defined cell-adhesive regions on an otherwise non-adhesive surface. Cells selectively attach only to these predefined areas, resulting in reproducible arrangements such as isolated single cells, clusters, or aligned structures. With ibidi µ-Patterning, this is achieved using the ibiTreat surface as the adhesive region, while the surrounding Bioinert surface prevents unwanted cell attachment. This creates ready-to-use micropatterned culture surfaces where cells attach only where intended.
What are the applications of micropatterning?
Micropatterning is used in many areas of cell biology and bioengineering where spatial control of cells is important. Typical applications include cell migration studies, cytoskeletal organization, mechanobiology, stem cell research, cell polarity studies, controlled cell–cell interactions, differentiation assays, and standardized spheroid generation. It is especially useful for microscopy-based experiments that require reproducible cell positioning.
What is the difference between single-cell, multi-cell, and line arrays?
Single-cell arrays are designed to position individual cells in isolated defined areas, making them ideal for single-cell analysis and morphology studies. Multi-cell arrays allow small groups of cells to attach together, which is useful for controlled cell–cell interaction studies or spheroid generation. Line arrays guide cells into elongated arrangements, making them useful for migration studies, cell alignment experiments, and polarization research.
How can micropatterning support spheroid generation?
Multi-cell arrays can be used to promote controlled cell aggregation and reproducible spheroid formation. By controlling where cells attach and interact, micropatterning helps generate more uniform spheroids compared to spontaneous aggregation methods.
Are ibidi µ-Patterning products suitable for live cell imaging and fluorescence microscopy?
Yes. ibidi µ-Patterning products are designed for microscopy-based workflows and are compatible with live cell imaging, fluorescence microscopy, and immunofluorescence staining. Defined cell positioning simplifies tracking and image analysis, while the ibidi Polymer Coverslip supports high-resolution imaging. For long-term live cell experiments, ibidi Stage Top Incubators help maintain stable environmental conditions directly on the microscope.
How does micropatterning improve experimental reproducibility?
One of the main advantages of micropatterning is improved reproducibility. Because cell position, shape, and spacing are controlled, experiments become more standardized and less dependent on random cell behavior. This is especially helpful in quantitative microscopy and comparative studies where subtle biological differences need to be measured reliably.
Are custom micropatterns possible at ibidi?
Yes. If standard pattern geometries do not match your experimental needs, custom micropatterns can be developed. ibidi offers customized µ-Patterning solutions with tailored shapes, dimensions, and layouts depending on the application. This allows researchers to create experimental designs optimized for specific cell types, workflows, or imaging requirements.