Chemotaxis Assays: Principles, Cell Migration Types, and Chamber Systems
What Is Chemotaxis in Cell Migration?
Chemotaxis is the directed migration of cells towards a chemoattractant and a key mechanism of cell migration in many biological processes. This process is different from chemokinesis, which is undirected cell migration.
Chemotaxis assays are widely used to study directed cell migration in vitro under defined gradient conditions. When cells undergo chemokinesis, they alter their migratory properties, for example by increasing or decreasing their migration speed due to a substance, but do not preferentially migrate in the direction of that substance.
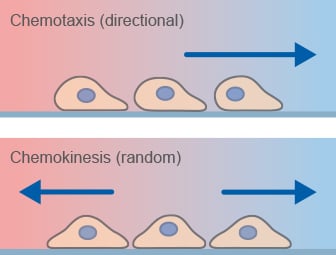
Chemotaxis-related migratory responses.
What is a chemotaxis assay?
A chemotaxis assay is a cell migration assay used to measure the directed movement of cells in response to a chemoattractant gradient.
What is the difference between chemotaxis and chemokinesis?
Chemotaxis describes directional cell migration along a gradient, whereas chemokinesis refers to non-directional changes in cell motility.
A chemotaxis assay is a specialized cell migration assay conducted to analyze whether or not a cell type directly orients and migrates towards a defined chemoattractant. Since chemotaxis is a particularly important process during cancer development and immune infiltration, frequently used cell types in chemotaxis assays are cancer cells and immune cells.
Chemotaxis assays enable the investigation of:
- chemotactic behavior of cells after knockout, knockdown, or overexpression of the gene of interest
- chemotactic potential of substances
- effect of chemotaxis inhibitors and enhancers
- differentiation between chemotaxis and chemokinesis
- cellular signal transduction during chemotaxis
- changes in the cytoskeleton during chemotaxis
These assays are widely used to study directed cell migration under controlled conditions in vitro.
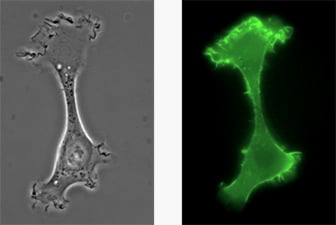
A chemotactic HT-1080 LifeAct-TagGFP2 cell migrating on a 2D surface.
Chemotaxis is induced by specific substances called chemoattractants. These substances can be a chemokine, a chemokine receptor, a growth factor, or a growth factor receptor. Important and well-described chemoattractants are, for example, the chemokine receptor CXCR4 and its ligand CXCL12, EGFR and EGF, and also CCR7 and CCL21.
The chemotactic network plays an important role in health and disease. On the one hand, chemotaxis is crucial in many physiological processes, such as during the recruitment of inflammatory cells or organ development. On the other hand, chemotactic processes can promote cancer progression, such as during the metastatic process, when malignant reprogramming of chemotaxis leads to cell invasion and dissemination.
Live cell imaging of actin dynamics in a LifeAct-expressing primary dendritic mouse cell after the application of a chemotactic gradient.
Tumor cells and their microenvironment constantly act in a complex communication network. Factors produced by epithelial tumor cells stimulate various immune cells, such as tumor-associated macrophages (TAMs), neutrophils, lymphocytes, and cancer-associated fibroblasts (CAFs). Upon stimulation, immune cells migrate towards the gradient, which can result in a tumor-suppressing or tumor-promoting immune response.
Chemotaxis is also strongly involved in angiogenesis—an essential process that occurs during organ development and cancer progression. Angiogenesis helps to supply the tumor with oxygen and nutrients and is required for tumor dissemination. Tumor cells produce chemokines that influence the migration of endothelial cells, resulting in the formation of blood vessels. Furthermore, chemokines can attract angiogenesis-promoting immune cells.
These complex biological processes make chemotaxis assays an essential tool to study cell migration under controlled conditions in vitro.
Types of Chemotactic Cell Migration
Chemotactic cell migration occurs in different modes depending on the cell type and microenvironment. These migration modes differ in speed, cell morphology, and interaction with the extracellular matrix.
Multicellular Migration
Collective Migration
Cells migrate as a coordinated group while maintaining cell–cell junctions (e.g., endothelial cells).
 |
|
Cell Streaming
Cells migrate individually but follow each other along defined paths.
 |
|
Single Cell Migration
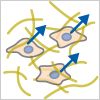
Amoeboid Migration
A fast migration mode characterized by low adhesion and high contractility.
 |
|
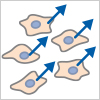
Mesenchymal Migration
A slower migration mode involving strong adhesion and ECM remodeling.
 |
|
Trepat X, Chen Z, Jacobson K. Cell migration. Compr Physiol. 2012;2(4):2369-2392. doi:10.1002/cphy.c110012.
Read article
Roussos ET, Condeelis JS, Patsialou A. Chemotaxis in cancer. Nat Rev Cancer. 2011;11(8):573-587. Published 2011 Jul 22. doi:10.1038/nrc3078.
Read article
Many cells naturally grow in a three-dimensional environment. When cultured in vitro, cells are attached to a flat 2D surface and might behave differently than when they are inside a 3D gel matrix. In many cases, a 3D environment more closely resembles an in vivo situation, and this should be considered when planning chemotaxis assays.
Using the µ-Slide Chemotaxis, chemotactic gradients can easily be established in water-based gels, such as Collagen I gels and Matrigel, because the gel structure does not hinder the diffusion. Learn more about the advantages and limitations of 3D chemotaxis assays here.
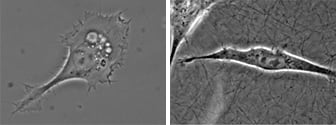
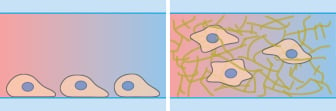
Microscopy and schematic of adherent HT-1080 cancer cells on a 2D surface (left), and embedded into a 3D Collagen I gel (right).
Chamber Systems for Chemotaxis Assays
Several chambers with different characteristics have been developed to perform chemotaxis assays, including the µ-Slide Chemotaxis, Boyden Chamber, Zigmond Chamber, and Dunn Chamber. These chemotaxis assay systems differ in gradient stability, imaging compatibility, and suitability for endpoint or live-cell analysis.
For detailed and reproducible analysis of chemotactic cell migration under stable gradient conditions, live cell imaging-based systems provide clear advantages over endpoint assays. The following comparison highlights the key differences between commonly used chemotaxis assay systems and helps identify the most suitable approach for your experiment.
- µ-Slide Chemotaxis – stable gradients and real-time cell tracking
- Boyden Chamber – endpoint migration assay (transwell)
- Zigmond Chamber – short-term gradient analysis
- Dunn Chamber – short-term chemotaxis studies
Comparison of Chemotaxis Assay Systems
| Feature | µ-Slide Chemotaxis | Boyden Chamber (Transwell) | Zigmond Chamber | Dunn Chamber |
|---|---|---|---|---|
| Assay Type | Live cell imaging | Endpoint assay | Live cell imaging | Live cell imaging |
| Gradient Stability | Stable (>48 h) | Steep, unstable | Short-term | Short-term |
| Cell Tracking | Yes (single-cell resolution) | No | Limited | Limited |
| 2D / 3D Compatibility | 2D and 3D | Mainly 2D | 2D | 2D |
| Quantitative Analysis | High precision | Limited | Moderate | Moderate |
| Cell Distribution at Start | Homogeneous | Inhomogeneous | Homogeneous | Homogeneous |
| Typical Use Case | Detailed migration analysis, live imaging, reproducible assays | Migration screening | Short-term gradient studies | Short-term gradient studies |
Among these systems, live cell imaging-based assays with stable gradients enable the most detailed and reproducible analysis of chemotactic cell migration. The µ-Slide Chemotaxis combines stable gradients with live-cell imaging, enabling precise and reproducible analysis of directed cell migration at single-cell resolution over extended time periods.
The µ-Slide Chemotaxis is a chemotaxis assay system designed to investigate the chemotactic behavior of fast or slow migrating, non-adherent or adherent cells on 2D surfaces or in 3D gel matrices. It is possible to observe cell migration in linear and stable concentration profiles for more than 48 hours. As gradients are rapidly established, fast migration responses (occurring in less than 30 minutes) can also be measured. This enables precise, reproducible analysis of chemotactic cell migration in real time.
- Cells migrate and can be observed in a central channel, which connects two large reservoirs
- Defined linear gradient with long-term stability
- Homogeneous cell distribution at the experimental starting point
- Ideal for live cell imaging using inverted microscopy
- Suitable for 2D and 3D assays
- Suitable for fast and slow migrating cells
- Suitable for adherent and non-adherent cells
Using the µ-Slide Chemotaxis allows for detailed and defined analysis of the migration behavior of various cell types, such as endothelial cells, fibroblasts, cancer cells, and immune cells.
Zengel P, et al. (2011) µ-Slide Chemotaxis: a new chamber for long-term chemotaxis studies. BMC Cell Biol 12:21. 10.1186/1471-2121-12-21.
Read article
Biswenger V, et al. (2018) Characterization of EGF-guided MDA-MB-231 cell chemotaxis in vitro using a physiological and highly sensitive assay system. PLoS One 13(9):e0203040.
Read article
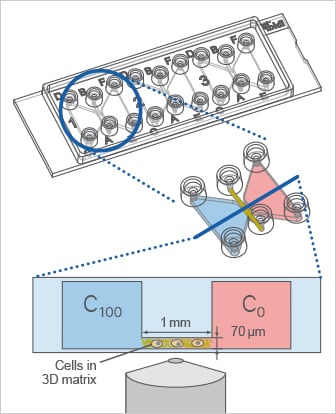
Using the µ-Slide Chemotaxis, the cells can be cultivated in a 3D gel matrix or on a 2D surface. In this example, a 3D experiment with migratory cells in a gel matrix is shown.
Gradient Stability in the µ-Slide Chemotaxis
The µ-Slide Chemotaxis is designed to establish a defined chemoattractant gradient with long-term stability for more than 48 hours. The gradient forms during the first few hours after adding the chemoattractant. After equilibrium is reached, it remains stable for the next 48 hours and then slowly flattens due to molecule transport across the observation area.
In 3D chemotaxis assays using aqueous gels such as Collagen I or Matrigel, diffusion of typical chemoattractant molecules is usually not impaired, because the gel pore sizes are large enough for molecular diffusion. If the matrix pore size is smaller than the diffusing molecule, however, diffusion can be reduced and gradient formation may be affected.
The Boyden chamber (transwell migration assay) is a widely used endpoint assay to quantify cell migration across a porous membrane but does not allow real-time tracking of individual cell behavior.
- Cells migrate from one side to the other of a filter or a porous membrane, also called transwell cell migration assay
- Endpoint assay: when analyzing a transwell migration assay, migrating cells are counted on the back side of the membrane, and therefore cell paths cannot be analyzed
- Steep gradients
- Inhomogeneous cell distribution at the experimental starting point; cells are situated on one membrane side only
Boyden S. The chemotactic effect of mixtures of antibody and antigen on polymorphonuclear leucocytes. J Exp Med. 1962;115(3):453-466. doi:10.1084/jem.115.3.453.
Read article
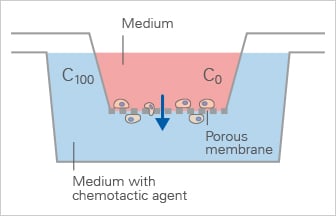
Schematic of the Boyden Chamber, in which cells migrate through a porous membrane towards a chemoattractant in an endpoint transwell migration assay.
The Zigmond chamber is an early chemotaxis assay system that allows observation of cell migration in a defined gradient but is limited by short-term gradient stability.
- Cells grow and migrate on a coverslip glass within a region of a bridge, which is located between two connected reservoirs
- Defined linear gradient
- No long-term stability
- Suitable for upright time-lapse microscopy
- Homogeneous cell distribution at the experimental starting point
Zigmond SH. Ability of polymorphonuclear leukocytes to orient in gradients of chemotactic factors. J Cell Biol. 1977;75(2 Pt 1):606-616. doi:10.1083/jcb.75.2.606.
Read article
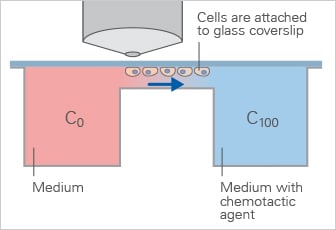
Schematic of the Zigmond Chamber, which generates a defined linear gradient across a bridge region for short-term chemotaxis experiments.
The Dunn chamber is similar to the Zigmond chamber and enables observation of chemotactic migration under defined gradients, but is primarily suited for short-term experiments.
- Similar to the Zigmond chamber
- Defined linear gradient
- No long-term stability
- Suitable for upright time-lapse microscopy
- Homogeneous cell distribution at the experimental starting point
Zicha D, Dunn GA, Brown AF. A new direct-viewing chemotaxis chamber. J Cell Sci. 1991;99 ( Pt 4):769-775. doi:10.1242/jcs.99.4.769.
Read article
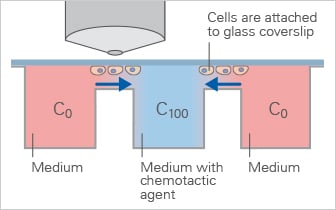
Schematic of the Dunn Chamber, a direct-viewing chemotaxis chamber used to study cell migration in defined gradients during short-term experiments.
Related Applications and Methods
Chemotaxis assays are closely linked to other cell-based workflows that help study cell migration, tissue-like environments, and dynamic cellular responses:
- Live cell imaging techniques for real-time analysis of cell migration dynamics
- 3D cell culture to study chemotaxis in physiologically relevant environments
- Angiogenesis assays to investigate endothelial cell migration and vessel formation
- Wound healing and migration assays to analyze collective cell migration and gap closure in 2D systems
Continue with the Chemotaxis Assay Workflow to learn how to set up chemotaxis experiments step by step, or explore Chemotaxis Data Analysis for key parameters, result interpretation, and example data.




