Chemotaxis Assay Workflow: From Sample Preparation to Data Analysis
A chemotaxis assay is a cell migration assay used to study directed cell migration along a defined chemical gradient. A reliable chemotaxis assay workflow includes experimental planning, sample preparation, chemoattractant gradient formation, live cell imaging, cell tracking, and quantitative data analysis.
This guide explains how to set up and analyze 2D and 3D chemotaxis assays using time-lapse microscopy. It covers key experimental parameters such as cell type, migration speed, imaging interval, assay duration, controls, and the number of cells required for meaningful single-cell tracking.
For reproducible chemotaxis experiments, the µ-Slide Chemotaxis for 2D and 3D chemotaxis assays enables stable gradient formation in parallel chambers and supports both slow- and fast-migrating cell types.
The workflow should include a negative control without chemoattractant (-/-), a positive control with uniform chemoattractant exposure (+/+), and an experimental gradient condition (+/-). These controls help distinguish directed chemotaxis from random migration and chemokinesis.
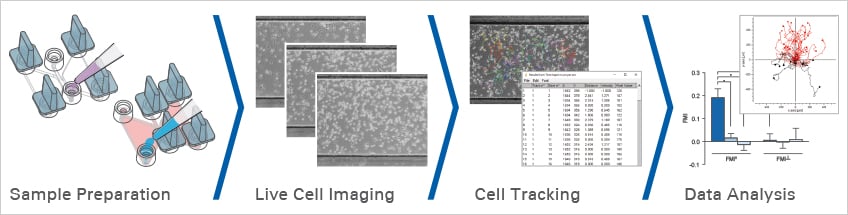
For a successful chemotaxis experiment, please address the key experimental questions before starting.
Briefly, define the cell type, migration speed, assay format, chemoattractant concentration, medium composition, imaging interval, experiment duration, number of biological repeats, and required controls.
Basic equipment requirements
- Inverted phase contrast microscope (5x or 10x objective recommended)
- Camera for time-lapse image acquisition
- Stage top incubator for long-term live cell imaging (required for most mammalian cell types)
Recommended extensions
- Motorized stage for parallel image acquisition
- Autofocus
Quick Tips for Chemotaxis Assay Setup
Microscope Setup: An inverted microscope is the most convenient setup for imaging chemotaxis assays using the µ-Slide Chemotaxis. Upright microscopes can also be used if the objective has a working distance of at least 14 mm.
Chemoattractant Concentration: The optimal chemoattractant concentration depends on the cell type, chemoattractant, and assay conditions. For initial experiments, use published concentrations whenever available. If no data are available, screen a concentration range in 10-fold steps, for example 1 nM, 10 nM, 100 nM, and 1 µM.
Positive Control Selection: For a positive chemotaxis control, use a chemoattractant that has been validated for your cell type. If no suitable reference is available, fetal calf serum can be used to induce a chemotactic response in many cell types.
Using the µ-Slide Chemotaxis for 2D and 3D chemotaxis assays, reproducible chemotaxis assays with defined chemotactic gradients can be performed.
First, the cells are seeded, which can be done either in a 2D or a 3D environment. After incubation and cell attachment, the two reservoirs of each chamber are filled with the chemoattractant, according to the determined loading scheme.
ibidi Solution
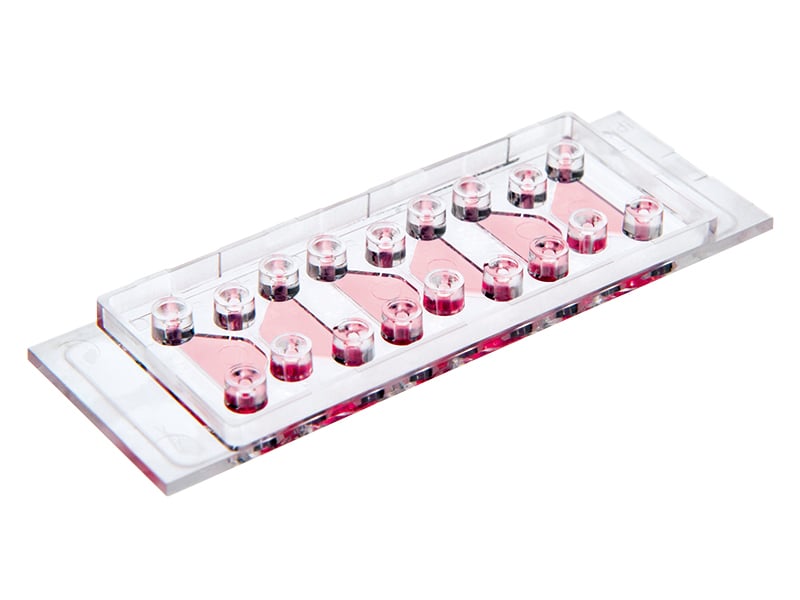

The µ-Slide Chemotaxis contains three separate chambers for parallel chemotaxis assays using slow or fast migrating cells.
It allows for the creation of a precisely defined, stable chemotactic gradient in a reproducible environment.
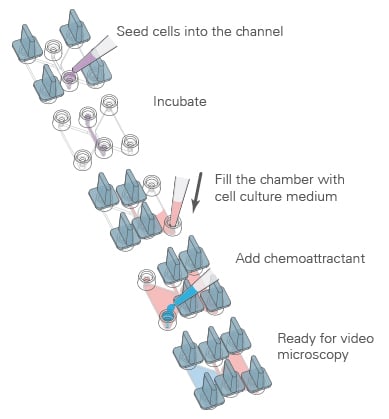
Live cell imaging under physiological conditions enables the documentation of cell migration over time and is essential for reliable chemotaxis assay analysis.
The duration of the imaging period depends on the cell type (e.g., fast migrating leukocytes or slow migrating tumor cells or fibroblasts) and the environmental conditions (e.g., type and concentration of the chemotactic agent). A typical video microscopy setup for slow migrating cells includes taking a photo every 2.5–10 minutes over a period of 24 hours.
Since each experiment optimally contains tracking data from 20–40 single cells, low-magnification microscopy objective lenses, such as 4x or 10x, should be used.
ibidi Solution

Maintain stable in vivo conditions during long-term chemotaxis imaging with ibidi Stage Top Incubators. They provide a physiological environment under the microscope, which enables live cell imaging during short-term and long-term chemotaxis assays.
Time-lapse microscopy of the migration of MDA-MB-231 human breast adenocarcinoma cells in a collagen gel. Cells were observed over a period of 24 hours in the µ-Slide Chemotaxis with EGF as chemotactic agent. Physiological conditions were maintained by an ibidi Stage Top Incubator.
Typically, live cell microscopy of a chemotaxis assay creates a temporal image stack, where each image displays the exact cell position at a specific time point.
For quantification of their movement, single cells are tracked over time by determining their position on each frame of the image stack. The tracking can be done either manually or automatically using special software. Afterwards, the positional values of each tracked cell (x, y) are available for each time point (t) and can be further analyzed.
Manual Tracking
The free ImageJ Manual Tracking Plugin assists in manually tracking cell migration in chemotaxis assays.
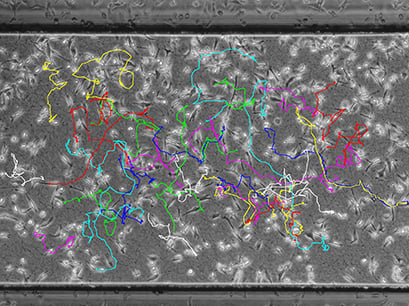

Visualization of cell traces after tracking with the ImageJ Manual Tracking plugin.
Read the detailed application chapter Chemotaxis Data Analysis: Parameters, Interpretation, and Example Chemotaxis Results to learn how to evaluate directed cell migration using trajectory plots, quantitative parameters, and statistical testing.
Explore the key steps of chemotaxis assay analysis:
Key Experimental Questions Before Starting an Experiment
In order to set up a chemotaxis assay correctly, it is crucial to answer the following questions first:
What is the migration speed/velocity of the cell type being analyzed?
Knowing your cell type of interest—including the culture medium requirements and the migration speed—is essential for subsequent assay planning. Whereas some cells have a low migration speed (e.g., tumor cells or fibroblasts), other cell types (e.g., leukocytes) migrate very quickly. The cell migration speed will determine the duration of the experiment and the intervals required between the images. Also, the gradient stability must be high enough to fit the total duration of the experiment. The µ-Slide Chemotaxis is suitable for chemotaxis assays with both slow and fast migrating cells.
| Slow migrating cells | Fast migrating cells | |
| Migration type | Mesenchymal | Amoeboid |
| Examples | Endothelial cells, cancer cells, fibroblasts, stem cells | T cells, neutrophils, dendritic cells, Dictyostelium discoideum |
| Speed | One cell length per hour: ∼10 µm/h | One cell length per minute: ∼10 µm/min |
| Frame rate (time-lapse) | 1 image per 10 min | 1 image per 30 s |
| Typical experiment duration | 12–24 h | 15–30 min |
What is the optimal culture medium?
Most cell types are cultured in a medium supplied with fetal calf serum (FCS). FCS contains many factors (e.g., enzymes, hormones and growth factors) that can influence the cell migration, and therefore may affect the outcome of a chemotaxis assay. For example, the effects of a chemotactic agent on cell migration can be masked by FCS-induced effects. Gradual reduction of the FCS concentration in the medium, before starting the assay, is one way to overcome this issue. Furthermore, special, serum-free culture media without FCS have been developed. The optimal medium composition for each cell type of interest must be tested before starting the assay.
What is the recommended seeding density?
The optimal seeding density depends on the cell type, proliferation rate, migration behavior, assay duration, and whether the cells are adherent or non-adherent. For adherent cells, about 200 cells should be present in the observation area of the µ-Slide Chemotaxis when starting live cell imaging. For non-adherent cells embedded in a gel matrix, a typical starting point is 6 µl cell suspension per chamber at 3 × 106 cells/ml, corresponding to approximately 18,000 cells per chamber.
For detailed protocols, refer to the chemotaxis Application Notes.
How many experiments should be performed?
Typically, three to five repeat experiments are sufficient to create significant data from the chemotaxis group and the respective control group. Each experiment should contain tracking data from 20–40 single cells, which is possible using low-magnification microscopy objective lenses, such as 5x or 10x.
Which controls must be included in the experiment?
For the correct analysis of a chemotaxis experiment, it is crucial to include a negative control without any chemoattractant (-/-), as well as a positive control with uniform chemoattractant exposure throughout the chamber (+/+).
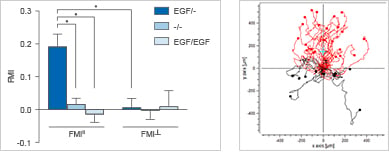
Using the µ-Slide Chemotaxis, a minimum of control measurements are required, since all conditions other than the gradient are symmetric. This allows for the analysis of chemotaxis and chemokinesis independently of each other. In the example, the chemoattractant induces both the chemotaxis and chemokinesis of cancer cells.
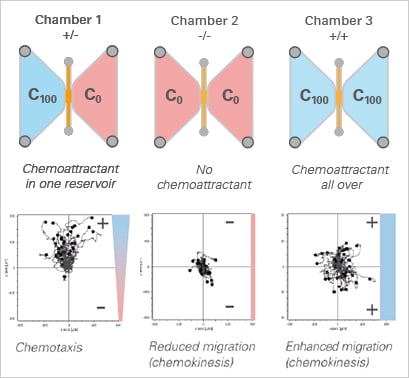
Experimental setup (top) and data plots (bottom) of a chemotaxis assay including the experimental group (+/-), a positive (+/+), and a negative (-/-) control group.
Should I perform a 2D or a 3D chemotaxis assay?
Most cells are naturally embedded in a 3D matrix. Culturing them in a 2D environment during a chemotaxis assay might alter their behavior and migrational capabilities. To overcome this issue, cells can be embedded in a 3D matrix that mimics their natural environment, such as collagen, Matrigel, or other hydrogels. The µ-Slide Chemotaxis is ideally suitable for both 2D and 3D experiments.
| Decision Criteria | 2D Chemotaxis Assay | 3D Chemotaxis Assay |
| Cell types | Cells that migrate on flat surfaces, such as adherent tumor cells, fibroblasts, or endothelial cells | Cells that migrate within extracellular matrices or tissue-like environments, including embedded adherent cells and suspension cells |
| Main advantage | Easy sample preparation, straightforward imaging, and simpler cell tracking | Highly defined, more in vivo-like environment with matrix-dependent cell migration |
| Biological relevance | Useful for standardized cell migration assays on defined 2D surfaces | Closer to physiological cell migration in extracellular matrix environments |
| Imaging and tracking | Simpler live cell imaging and 2D cell tracking | Cells may migrate in different focal planes and may go out of focus at higher magnifications |
| Limitations | Less physiological for cells that naturally migrate in 3D matrices | More demanding gel handling, additional assay parameters, and possible focus changes during imaging; cells might attach to 2D surface, thus creating 2.5D conditions |
| Recommended when | You need a robust, easy-to-handle chemotaxis assay with clear 2D migration readout | You want to study chemotaxis in a more tissue-like, matrix-based environment |
Find more information about 2D and 3D chemotaxis assays in the following Application Notes:
Read more about chemotaxis in cell physiology and chemotaxis assay methods, or learn about chemotaxis data analysis and interpretation.
Chemotaxis Assay Troubleshooting
My cells do not migrate. What can I do?
Cell migration is highly sensitive to substrate, coating, medium composition, supplements, cell density, passage number, and cell state. If cells migrate very slowly or do not migrate at all, test different experimental conditions while changing only one parameter at a time. Some cell types may also require specific stimulation or differentiation before they become migratory.
How can I avoid bubbles in the µ-Slide Chemotaxis?
Air bubbles often occur when gas is released from the medium or polymer after a temperature increase. To reduce bubble formation, degas all liquids and the µ-Slide Chemotaxis by placing them in the incubator for 24 hours before starting the experiment.
How can I improve gel loading?
Gel loading can be sensitive to pipetting technique, gel viscosity, temperature, and handling time. Work carefully, avoid introducing air, and follow the detailed troubleshooting recommendations in the µ-Slide Chemotaxis Instructions.
Frequently Asked Questions on Chemotaxis Assay Workflows
Which chemoattractant concentration should be used?
The optimal concentration depends on the cell type, chemoattractant, and assay setup. Start with published concentrations for your cell type if available. If not, screen a concentration range in 10-fold steps, for example 1 nM, 10 nM, 100 nM, and 1 µM.
Which controls are needed for a chemotaxis assay?
A chemotaxis assay should include a negative control without chemoattractant (-/-), a positive control with uniform chemoattractant (+/+), and the experimental gradient condition (+/-). The controls help distinguish directed chemotaxis from random migration and chemokinesis.
How long should a chemotaxis assay be imaged?
As a rough guideline, cells should be followed until they have migrated at least 10 times their cell diameter. The exact imaging duration depends on the cell type and migration speed. Slow migrating cells are often imaged for 12–24 hours, while fast migrating cells may require shorter experiments with higher frame rates.
How many cells should be tracked in a chemotaxis assay?
Usually, 20–40 cells per observation field are sufficient. For weak chemotaxis effects, more cells may be needed to reach statistical significance. For strong chemotaxis effects, fewer than 20 cells per condition may be sufficient.
Should I use a 2D or 3D chemotaxis assay?
The choice depends on the biological question and cell type. A 2D chemotaxis assay is easier to handle and image, while a 3D chemotaxis assay provides a more in vivo-like environment for cells that naturally migrate within extracellular matrices such as collagen or other hydrogels.




